|
12/3/2023 0 Comments Clc genomics workbench downloadFixed: Motif search did not exclude regions with Ns when the option 'Exclude matches in N-regions for simple motifs' was selected.Fixed: importing adapters for trimming and barcodes for de-multiplexing did not work properly for CSV files and empty rows in Excel files were not allowed. 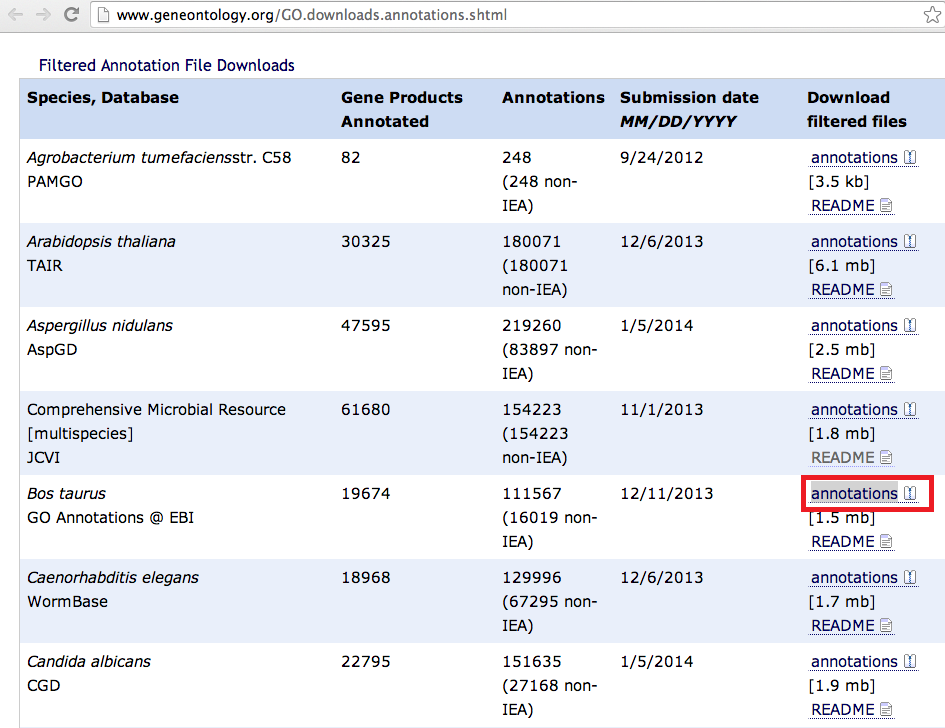
We recommend checking your mapping results manually if you rely on using the consensus sequence for further analysis. High gap counts would then kick in further downstream, possibly making the consensus a gap where it should not be. From that point onwards, gap counts are included in the consensus vote, but they are taken from the start of the mapping (where they are all 0), so they are out of sync with associated base counts. Note If you wish to import the reads in a SAM/BAM file as a sequence list, disregarding any mapping information, please use the Standard import instead. It occurs when 1) there is no coverage in an initial segment of the reference sequence, and 2) a gap is encountered in the global read alignment. The CLC Genomics Workbench includes support for importing SAM and BAM files from Complete Genomics. CLC Genomics Workbench and CLC Genomics Server packages provide tools for next generation sequence. Fixed: Calculation of consensus sequence in read mappings: Sometimes a majority of gaps would be ignored and a base erroneously introduced in the consensus sequence. MatLab is also available for download through UDeploy.Commonly, this program's installer has the following filenames. The most popular versions among CLC Main Workbench users are 7.6, 7.5 and 7.0. The actual developer of the software is CLC bio, a QIAGEN Company. CLC Genomics Workbench 23download premium module full crack with license is a powerful software developed by QIAGEN for scientists to analyze and visualize. The software is categorized as Photo & Graphics Tools. The annotation was only displayed on the first line. Our website provides a free download of CLC Main Workbench 8.1.2. Fixed: Annotations spanning the sequence from start to end did not display right when the sequence was wrapped.Exporting fastq format no longer includes redundant name of the read in the quality score line.They always applied to the first sequence although the fragment was located on another sequence (as indicated in the table). Fixed: When using multiple template sequences, the choices to open or annotate a fragment from the fragment table did not work properly.a T would still be a mismatch if the template sequence has an R, which means either A or G). Note that the primer base of course need to be covered by the ambiguity symbol (e.g. 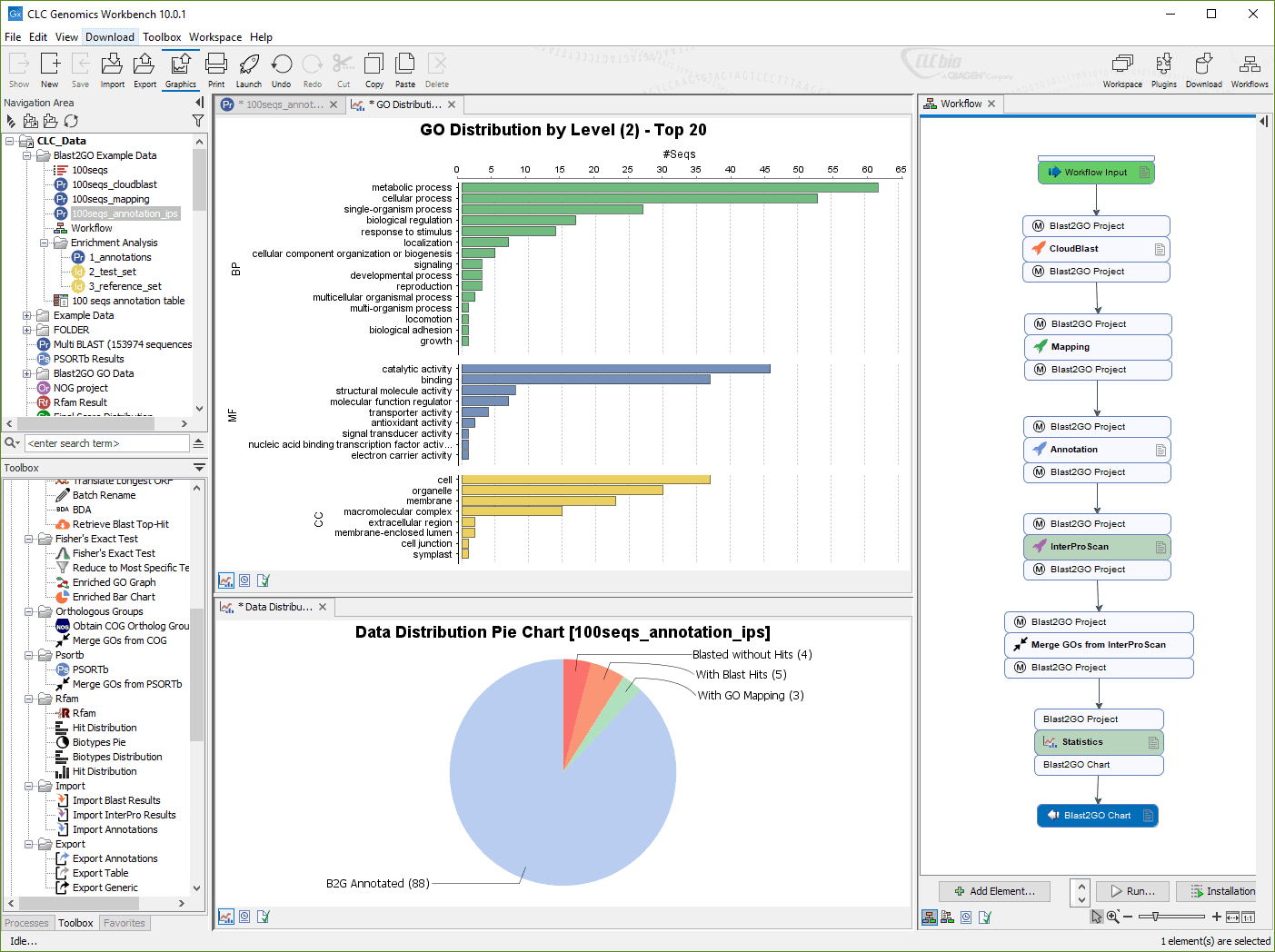 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
